Doctor in the Loop
Improve learning efficiency by closing the loop
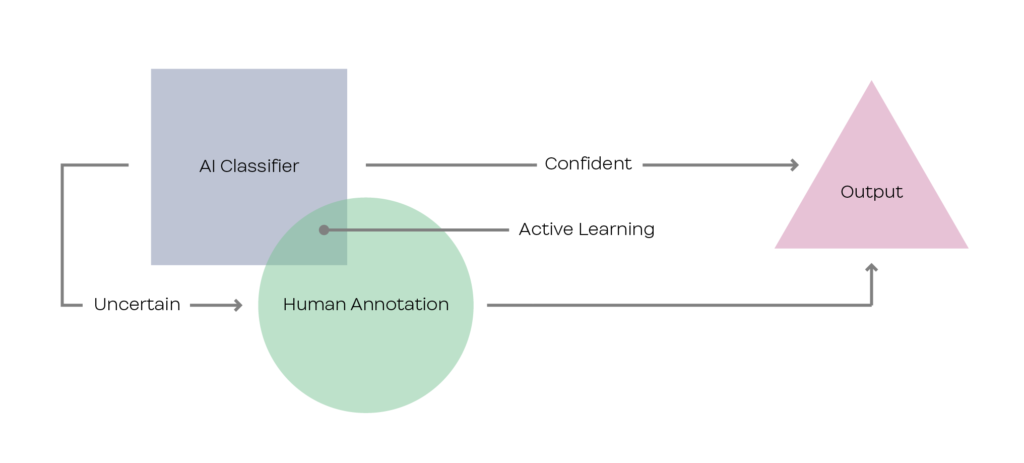
Background
In the past decade, despite showing promising success in many aspects of biomedical applications such as semantic segmentation of biomedical images [1], machine learning (ML) methods in healthcare is often limited by the cost of getting clinicians to annotate a large number of samples. In the context of ophthalmology and inherited retinal diseases (IRDs) study, labelling areas of images associated with IRD can be extremely labour intensive and requires the expertise of experienced clinicians. To this effect, works in the field have explored ways to perform tasks with limited amounts of samples. One potential approach is to explore active learning with bayesian neural networks [2] using uncertainty measurements [3,4]. In this project, the student will explore active learning approaches for medical image segmentation in the context of ophthalmology and build upon established architectures [5,6,7].
Goals
- Quantify the accuracy of computer vision models in IRD image classification/segmentation when trained on randomly sampled records.
- Increase the label quality in the training dataset by actively inquiring about corner cases where the model is most uncertain
- Minimize annotation time by not inquiring about trivial cases (active learning), and by stopping the process when the desired accuracy is reached.
- Analyze, qualitatively and quantitatively, the sampling techniques with random sampling baseline on various existing image segmentation architectures
Work plan
- Get acquainted with the workspace, dataset, and workflow.
- Review state of the art research in human-in-the-loop learning, especially in the context of biomedical image analysis.
- Replicate and adapt these research to the Moorfields eye hospital dataset.
- Acquire annotations from clinicians on fortnightly basis.
- Clean up and write up.
Prerequisites
The ideal student should have a good understanding of probabilistic reasoning and bayesian statistics to be able to conceptualize the statistical guarantees and optimal sampling needed for active learning. It is highly recommended to have some understanding of and interest in computer vision and image segmentation methods. On the practical side, experience with python and some of its scientific computation packages such as numpy, scikit learn, and pandas is a must. Experience with deep learning packages such as TensorFlow, Keras or PyTorch is a bonus. We strongly advise students to have a good understanding of bash scripts as the project is conducted mostly on our Linux server.
References:
- Ronneberger, O., Fischer, P., and Brox, T.. U-Net: Convolutional Networks for Biomedical Image Segmentation.
- Budd, S., Robinson, E. C., & Kainz, B. (2021). A survey on active learning and human-in-the-loop deep learning for medical image analysis. Medical Image Analysis, 71, 102062. https://doi.org/10.1016/j.media.2021.102062
- Depeweg, S., Hernández-Lobato, J., Doshi-Velez, F., and Udluft, S.. (2017). Decomposition of Uncertainty in Bayesian Deep Learning for Efficient and Risk-sensitive Learning.
- Chen, X., & Zhang, N. (2021). Model performance inspection of deep neural networks by decomposing bayesian uncertainty estimates. 2021 International Joint Conference on Neural Networks (IJCNN). https://doi.org/10.1109/ijcnn52387.2021.9533764
- J. Long, E. Shelhamer, and T. Darrell. Fully convolutional networks for semantic segmentation. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2015.
- Badrinarayanan, V., Kendall, A., Cipolla, R.: Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Transactions on Pattern Analysis and Machine Intelligence 39(12), 2481–2495 (2017)
- Ferianc, M., Manocha, D., Fan, H., and Rodrigues, M.. (2021). ComBiNet: Compact Convolutional Bayesian Neural Network for Image Segmentation.



